#500 – #define and #undef Must Be at Top of File
January 18, 2012 Leave a comment
In C#, all #define and #undef preprocessor directives must appear at the top of a source code file, i.e. before any valid C# tokens. The definition (or un-definition) then remains in effect until the end of the file is reached.
#define QUIET
using System;
using DogLibrary;
namespace ConsoleApplication1
{
class Program
{
static void Main()
{
Dog d1 = new Dog("Kirby", 12);
#if QUIET
d1.Bark("Arf");
#else
d1.bark("WOOOF");
#endif
}
}
}
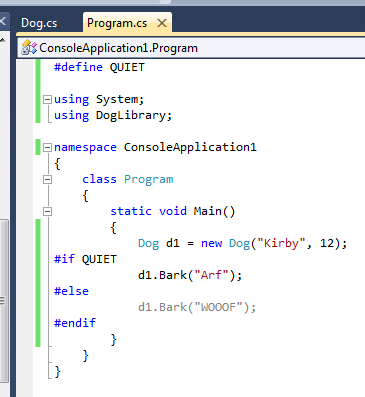
In this case, because the conditional compilation symbol is included directly in the code, Intellisense already knows which of the two Bark methods will be included in the code. It displays the line that will not be compiled in grey.
In this example, the second Bark method is not compiled at all.